Monoclonal antibodies have been effective in treating in both preventing and treating SARS-CoV-2 infections. However, just as SARS-CoV-2 evolves to evade immune responses by vaccines and natural infection, so too do variants arise that evade neutralization by specific monoclonal antibodies, hence the search for antibodies that broadly neutralize regardless of the source. We have previously described eight such antibodies derived from infected humans, mice, and camelids. Here we describe a different approach in which antibodies are isolated from monkey subjects.
In the search for such antibodies, He et al. discovered a series of antibody candidates induced in macaque monkeys. The researchers immunized macaques over a ten-week period and drew the antibodies from sera post-inoculation (Figure 1A). They used a recombinant prefusion-stabilized soluble S-protein plus a saponin (SMNP) adjuvant as their vaccine, yielding strong antibody responses in the animals.
Neutralization of SARS-CoV-1 and SARS-CoV-2
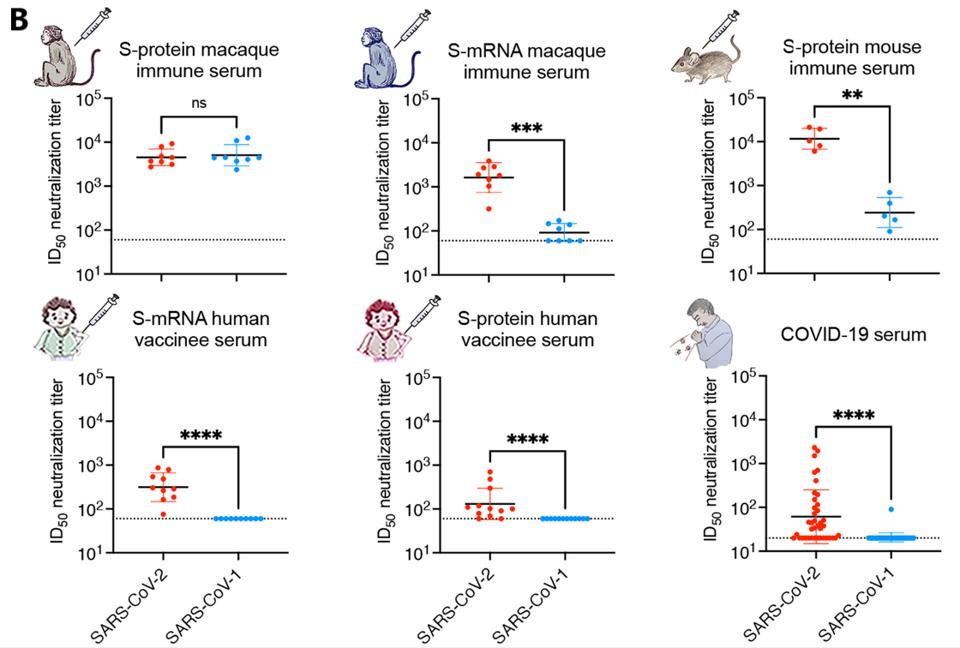
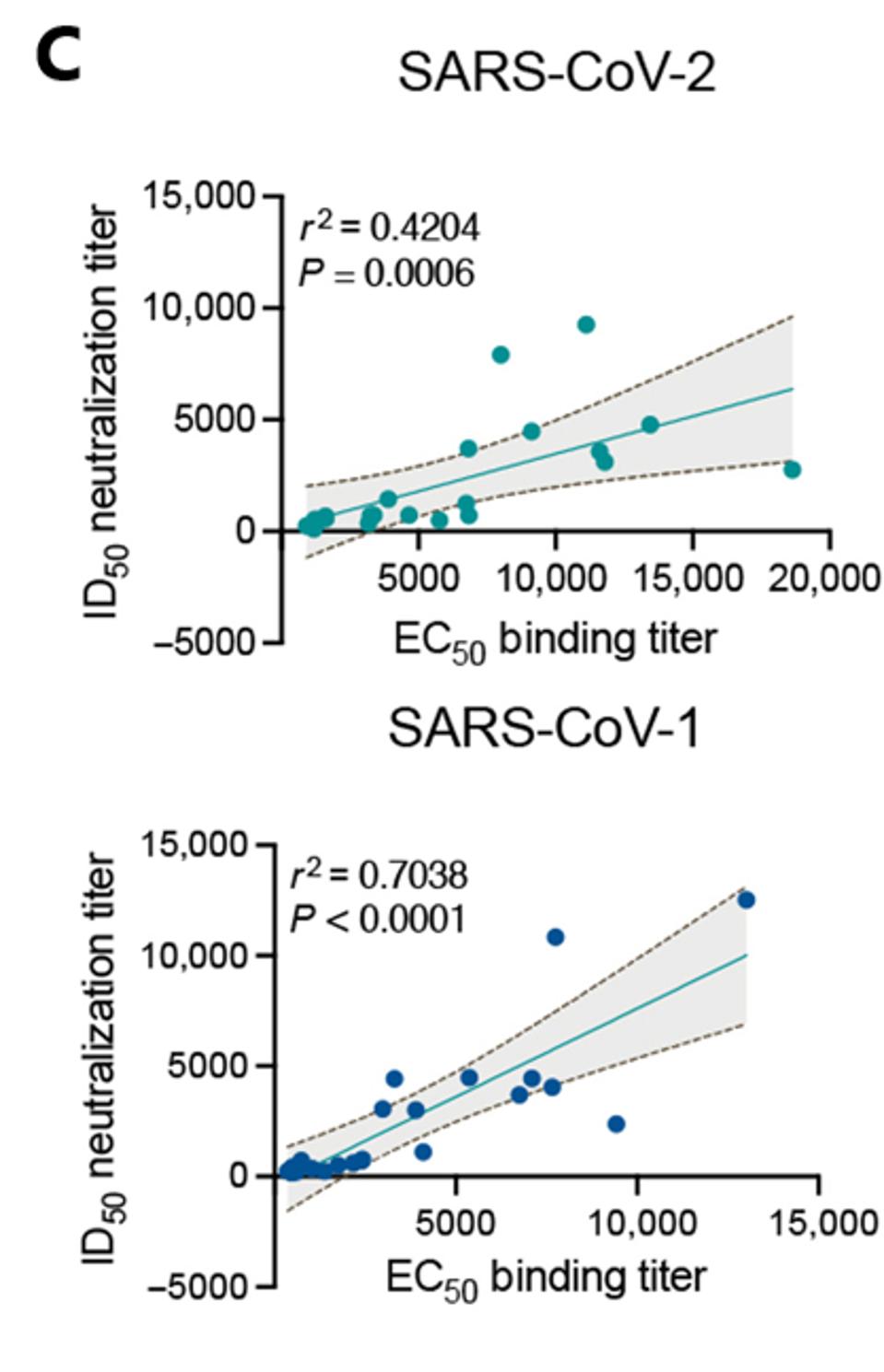
Not only did the enhanced adjuvant immune serum grant the macaques with significant virus neutralization of SARS-CoV-2, but the treatment also neutralized macaques infected with SARS-CoV-1, indicating broad neutralization capability (Figure 1B). This response is in stark contrast to human natural infection, which rarely induces cross-neutralizing activity against SARS-CoV-1. Inoculation with mRNA SARS-CoV-2 did not yield SARS-CoV-1 neutralization in monkeys or humans, and inoculation with the enhanced adjuvant S protein only yielded a significant SARS-CoV-1 response in monkeys. The sera of adjuvant vaccinated monkeys bound and neutralized both viruses with great consistency (Figure 1C).
Isolation of Monoclonal Antibodies
The intention behind using enhanced adjuvant S protein as a vaccine in monkeys is to enable enhanced memory B cell responses, from which strong antibody candidates can be isolated. These differ from most monoclonal antibody candidates, which derive from plasma cells.
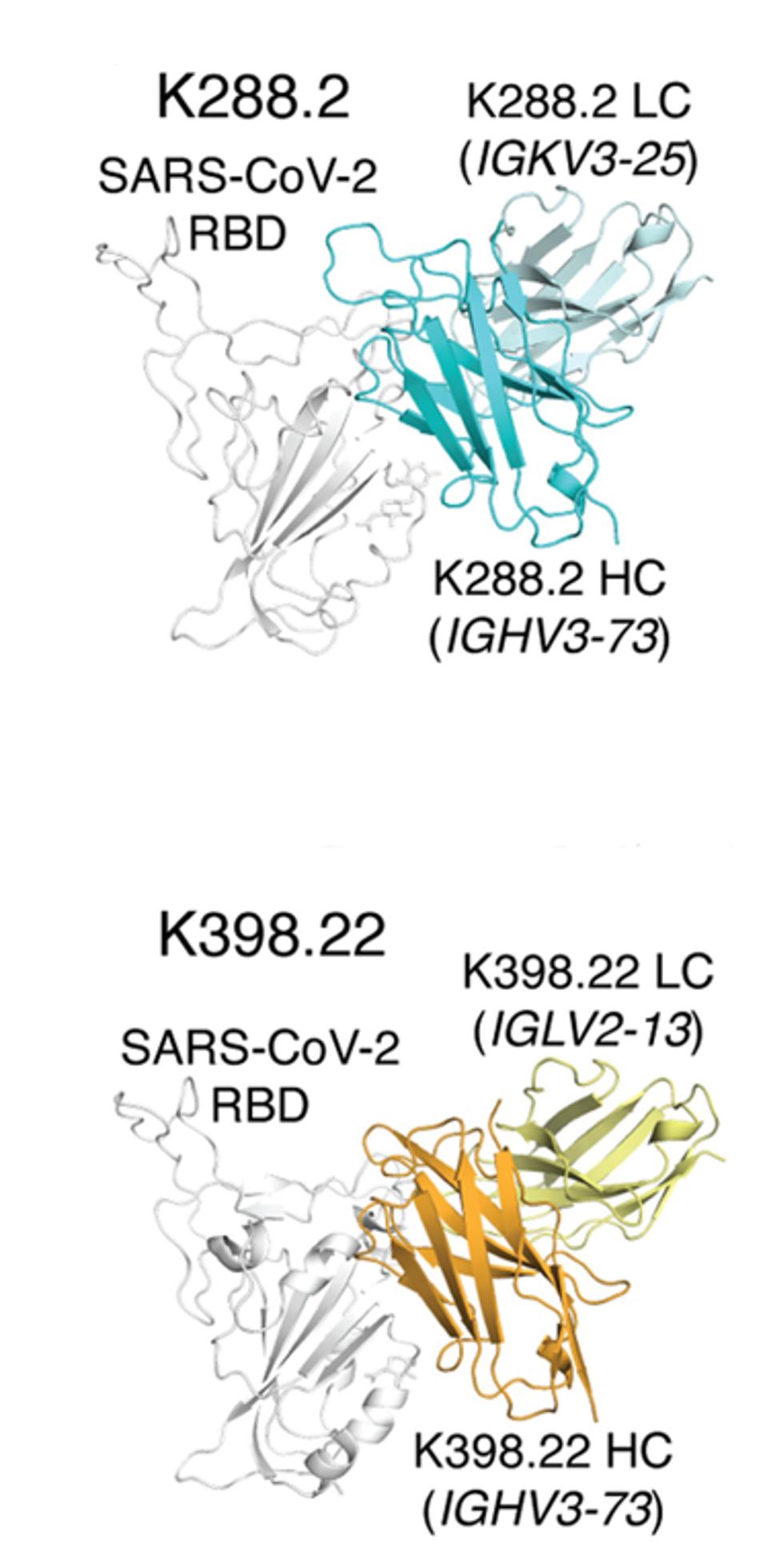
The researchers isolated many monoclonal antibody candidates from the macaque sera, but two stood out for binding and neutralization: K288.2 and K398.22. Both bind and neutralize a wide variety of viruses, including both SARS viruses and bat viruses WIV1 and RaTG13. Additionally, both neutralize variants of SARS-CoV-2 from the wildtype through Delta effectively. However, K288.2 loses neutralization capability against BA.1, whereas K398.22 maintains it. Figure 2 below illustrates both antibodies’ similar but slightly adjusted binding actions, which is likely the culprit for K288.2 losing neutralization against Omicron, binding to the wrong residue.
Cyro-Electron Microscopy: The Angle May Make All The Difference
The researchers note that differences in binding capacity between different viruses may be attributed to inherited differences between macaque and human sera from the immune system. Each species uses a different immunodominant germline gene segment in its antibody development, leading to different approach angles and binding modes when introduced to the receptor-binding domain of SARS-CoV-2 and others.

High-Resolution X-Ray Crystallography
X-ray crystallography allows researchers to observe specific amino acid binding residues as the antibodies come in contact with the virus. Not only do K288.2 and K398.22 bind conserved regions in the receptor-binding domain that are less often targeted in other monoclonal candidates, such as 373-375, 404-408, and 502-508, but there is additional evidence suggesting that the macaque germline introduces a specifically unique binding motif not found in human-derived antibodies, E33 in CDRH1, which perhaps is another cause for the bolstered SARS-CoV-1 neutralization in the macaque antibodies.
Caveats
The first caveats come from the isolation of the antibodies themselves. The sera was developed from an irregular and powerful adjuvant that was administered over a much longer 10-week period, meaning comparison to mRNA-derived sera in a more consistent four-week period is not directly apples to apples.
These antibodies would be of much greater significance if they demonstrated binding and neutralization of later Omicron variants such as BA.5, BA.2.75, or BA.4.6, though there have been no published results discussing these variants. Through personal communications with the authors, we have confirmed that K398.22 neutralization drops significantly against BA.2, which does not bode well for later Omicron variants.
Speculation
The macaque isolation process may be a favorable system for isolating new monoclonal antibodies to be used as drugs. The Omicron viruses circulating today have been selected against the human immune responses, not macaque. This study shows that the inherited germline sequences in macaque antibodies may give them a leg up against emerging variants tailored to evade human antibodies. That is not to say that K288.2 and K398.22 evade currently circulating strains. Rather, the method used to discover these two may yield others yet to come and the macaque isolation process is one that should be thoroughly investigated.