This is part of a continuing series describing antiviral antibodies to prevent and treat SARS-CoV-2 infections. In this series, we will discuss the fundamental nature of virus evolution, how SARS-CoV-2 has mutated to evade neutralizing antibodies, and our latest attempts to fight against these mutations with more recent and improved antibody candidates.
SARS-CoV-2 variants that evade the immune system are characteristic of the Covid-19 pandemic. As novel SARS-CoV-2 variants develop new mutations, their evasion from existing treatments and vaccines continues to increase. Antibodies from vaccines wane after a few months, antibodies from previous infections are often ineffective, and monoclonal antibody therapies that worked against earlier variants struggle against current variants. The current state of the pandemic has sparked a search for monoclonal antibodies that neutralize not one but all variants to counter these new variants.
In this series, we have discussed several pan-variant monoclonal antibodies, all of which promise against current Omicron variants and previous variants of concern, such as Alpha, Beta, and Delta. Here we analyze two new antibodies described in a study by Fenwick et al. that target a conserved site in SARS-CoV-2 variants.
Antibody Discovery
Collecting sera samples from over 100 donors of varying vaccine and infection status, Fenwick and colleagues focused their search on the sera of a post-infected donor who had received a full dose of Moderna mRNA vaccine, yielding six monoclonal antibodies for cloning and testing.
The six clones were introduced to a binding assay against a panel of SARS-CoV-2 variants to whittle further the field, including the original Wuhan Spike, Alpha, Beta, Gamma, and Delta. Two antibodies, P2G3 and P5C3, outperformed the five others and became the focal point of examination for Fenwick and colleagues moving forward.
Neutralization Comparison to Existing mAb Therapies
Currently available monoclonal antibody treatments, such as the AstraZeneca cocktail (cilgavimab + tixagevimab), the Regeneron cocktail (casirivimab + imdevimab), and Sotrovimab, are not effective against the Omicron family of variants, including BA.1, BA.2, BA.5, and BA.2.75, with the possible exception of bebtelovimab and sotrovimab against BA.2.75.
Not only does the P2G3 antibody bind and neutralize the Wuhan, Alpha, Beta, and Delta variants to a greater degree than any of the above-listed monoclonal treatments. It also wholly neutralizes live Omicron BA.1 and BA.2 virus. Both alone and in combination with P5C3, the P2G3 antibody’s IC50 values consistently remain under .025 µg/ml.
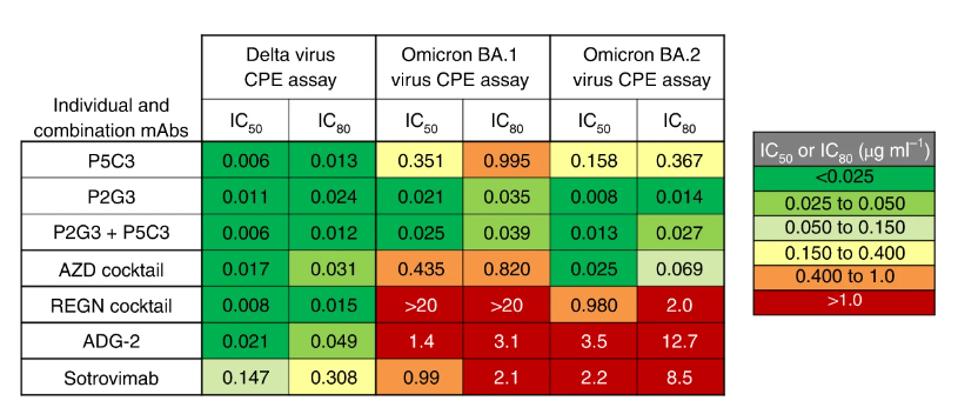
Notably, Fenwick and colleagues did not include neutralization data for the currently circulating BA.5 or BA.2.75 variants. However, they suggest that the novel mutations in those viruses would likely not impact neutralizing activity. Regardless, a further examination should be taken to confirm that hypothesis.
Antibody-Dependent Cellular Cytotoxicity and Antibody-Dependent Cellular Phagocytosis
The researchers next examined the antibodies for antibody-dependent cellular cytotoxicity (ADCC) and phagocytosis (ADCP). Both function to clear infected cells from the host. ADCC is the enabling of host immune cells to kill virus-infected cells, whereas ADCP is the enabling of host immune macrophages to absorb and kill infected cells.
They found that P2G3-mediated ADCC was more robust than any other monoclonal antibody tested, including the other cloned antibodies and the AstraZeneca and Regeneron cocktails. For ADCP, Fenwick and colleagues found greater activity in the P2G3 + P5C3 combination rather than in isolation, adding a significant data point in favor of the tandem.
Live Animal Studies
To examine the use of P2G3 as both a prophylactic and therapeutic in live hosts, Fenwick and colleagues administered the antibody to both hamster and macaque models. In hamsters, the antibody was administered two days before infection with the Wuhan virus. After four days of infection, only one of six infected hamsters had detectable infectious virus in the lungs, which was a massive four-log reduction from the control group.
In the macaques, the P2G3 antibody was administered 72 hours before infection with the BA.1 virus. The researchers found that peak viral load in the treated macaques was roughly half that of the control group, suggesting P2G3 maintains therapeutic efficacy even against later Omicron viruses.
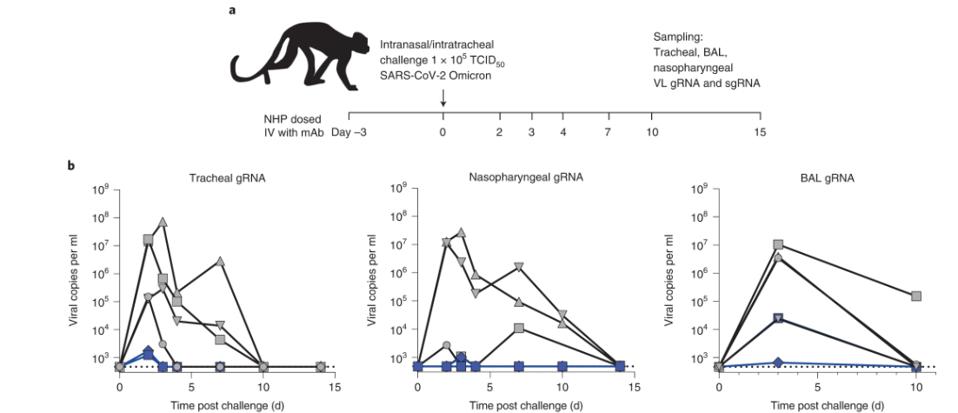
Binding Epitope
P2G3 and P5C3 fall into the antibody group that binds the receptor-binding domain, many of which we have described in this series.
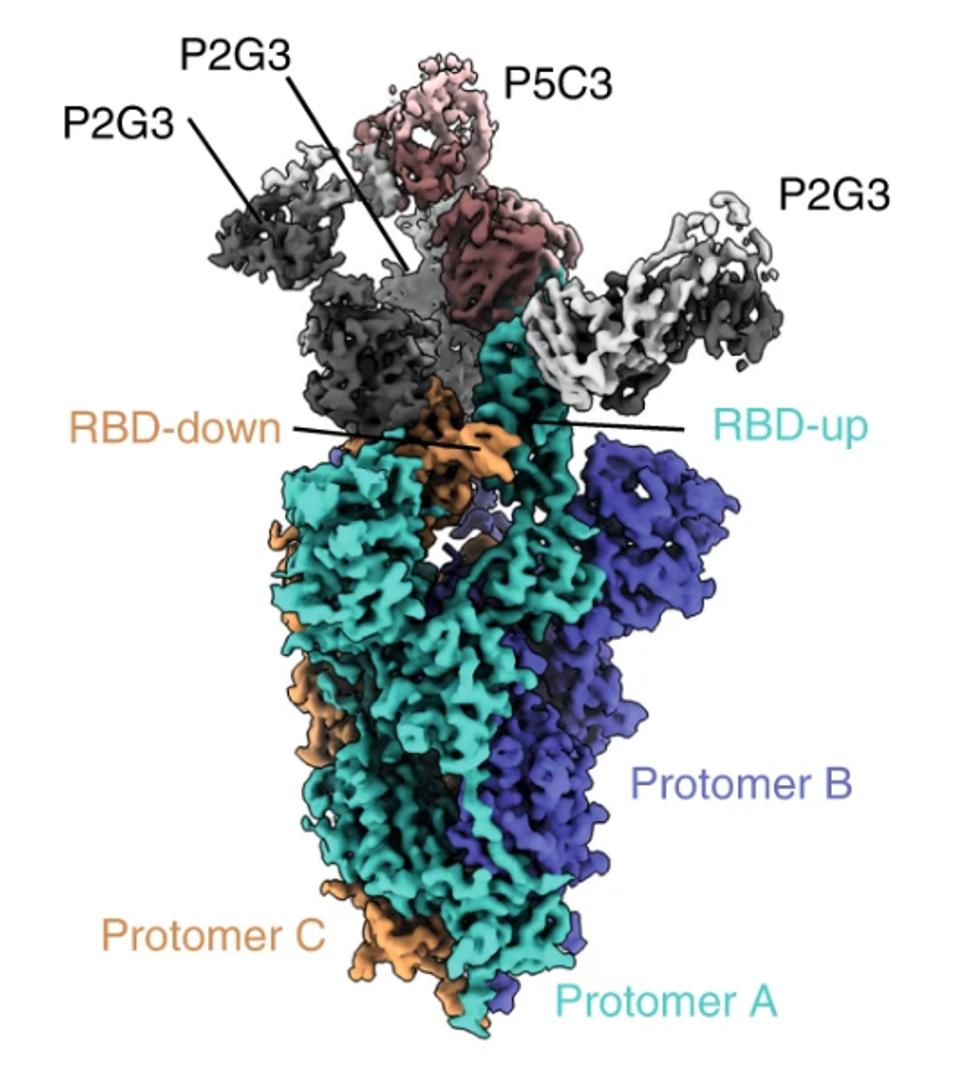
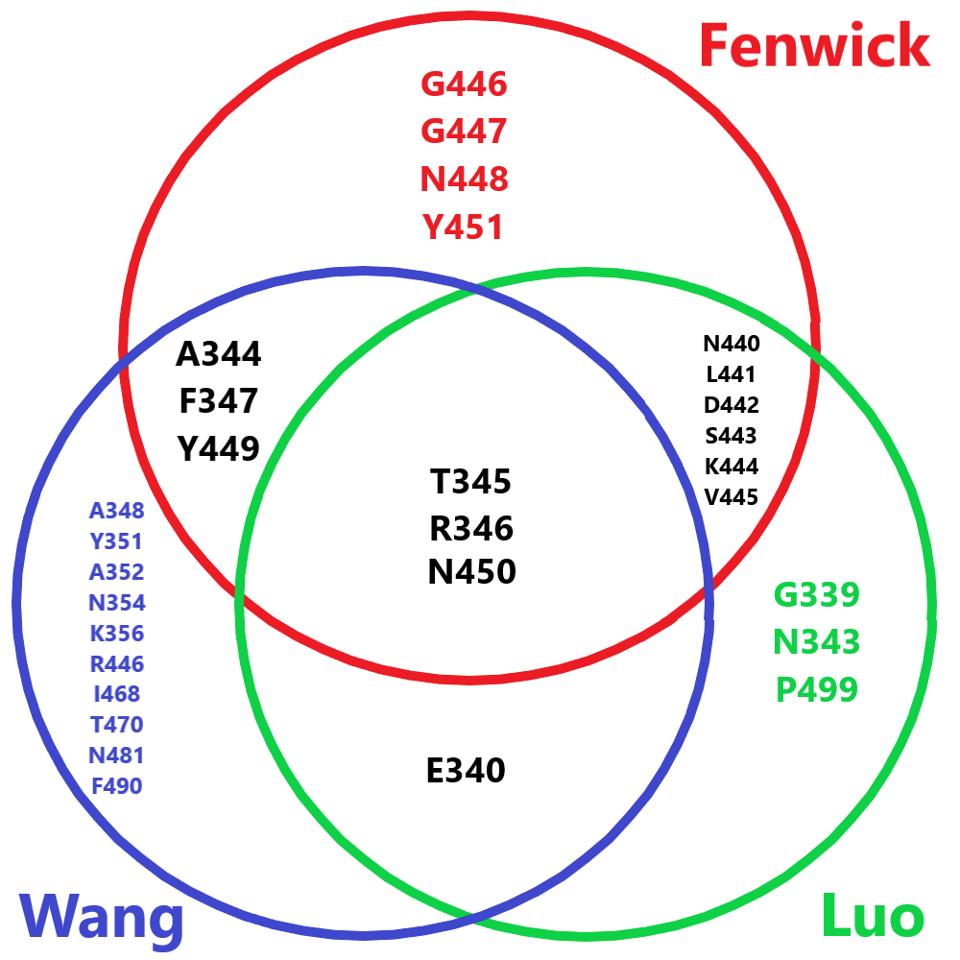
Using the widely adopted method of cryo-electron microscopy, Fenwick and co. found the exact binding epitope of the antibodies. The two antibodies in conjunction interact with residues 344-347 and 440-451 for a total of 16 receptor-binding domain contact points. According to the GISAID sequence database, these residues are highly conserved, aside from three notable exceptions, all commonly found in the major versions of the Omicron family. However, as the neutralization assays demonstrate, these mutations clearly do not impede Omicron neutralization.
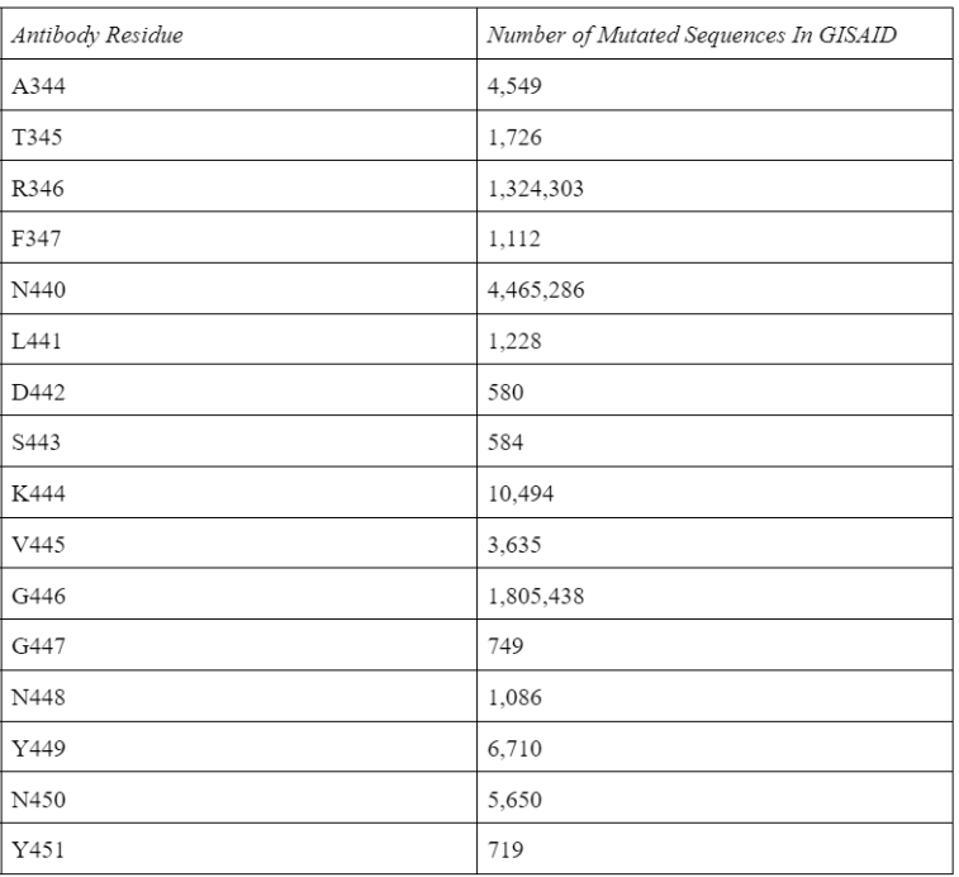
The Fenwick antibodies share many overlapping residues with antibodies we have previously described in this series. Most notably, the Luo and Wang antibodies. All three are viable receptor-binding domain targeting antibodies that would prove vital to combination therapy in tandem with fusion domain and S2 targeting antibodies.
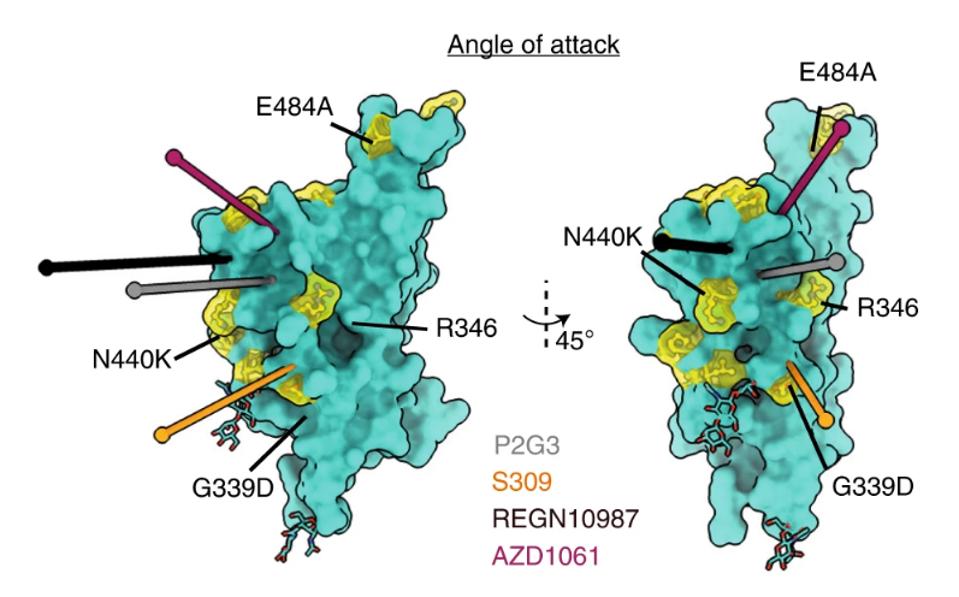
The researchers also make a note of P2G3’s angle of attack. Many monoclonal antibodies are structured such that they can only bind at certain angles to the receptor-binding domain when it is in a specific conformation, whether up or down. This includes the Regeneron cocktail REGN10987, which can only bind to the up-RBD. In contrast, P2G3 can bind in either conformation from a different angle, giving the antibody more flexibility and potentially one catalyst behind its broad neutralization.
Conclusion
While this antibody lacks data on the latest versions of Omicron sweeping the world, its binding and neutralization speak for themselves. It shares many binding residues with antibodies that we know are effective against the latest variants and outcompetes many commercially available monoclonal therapies. Add this antibody to the growing list of weapons we can use to combat the everpresent SARS-CoV-2 virus as Omicron continues to infect, mutate, and spread.


