The greatest difficulty for vaccine and drug design three years into the Covid-19 pandemic is the proliferation of viral variants. Most antibodies target the SARS-CoV-2 receptor-binding domain, but this region is highly variable, leading to variants of concern such as Beta, Delta, and now Omicron.
A potential solution to this problem is to find antibodies that target conserved sites unlikely to mutate. Many have attempted to locate these antibodies that target sites like this in the receptor-binding domain, though they are often still overwhelmed by the latest iterations of Omicron.
Researchers Zhou et al. from the Scripps Research Institute in La Jolla, California, examine a different approach. Their study analyzes a panel of broadly neutralizing antibodies that target not the receptor-binding domain but the S2 stem-helix fusion region of the spike. Zhou and his team previously isolated the CC40.8 antibody, which targets this conserved region of SARS-CoV-2, binding and neutralizing in mouse models. Notably, the antibody also protected against sister viruses SARS-CoV-1 and MERS-CoV, speaking to the broad capabilities of S2 binding antibodies. Here we examine their results and the impact on antibody development moving forward.
Study Design
Zhou et al. isolated antibodies from the sera of a wild variety of donors. 15 patients were Covid-19 recoverees, 10 had received two doses of mRNA vaccine, nine had received three doses, and 15 had recovered from Covid-19 and received one dose of mRNA vaccine.
Notably, none of the sera from the recoveree, two-dose, or three-dose groups bound to spike S2 stem-helix peptides. However, 80% (12/15) exhibited strong binding to these epitopes in the recovered-vaccinated group. Accordingly, Zhou et al. focused on this group from here, aiming to isolate antibodies for a larger panel examination.
From ten of the 12 recovered-vaccinated patients, Zhou et al. isolated 40 stem-helix monoclonal antibodies. 38 of the 40 bound a range of coronaviruses, including SARS-CoV-1, MERS-CoV, and SARS-CoV-2. Some antibodies were duplicates, reducing their pool to 32 broadly binding antibody candidates.
Broad Neutralization of Betacoronaviruses
They next introduced the panel of 32 antibodies to another host of viruses to determine neutralization efficiency. All 32 antibodies neutralize viruses such as SARS-1, MERS, and SARS-CoV-2 to varying degrees. When including MERS-CoV, only 72% (23/32) efficiently neutralized across the board.
Further narrowing their group, they selected the top ten highly potent antibodies against the previous group of viruses to introduce to a range of SARS-CoV-2 variants of concern. Consistent with their previous findings, all ten demonstrated highly effective neutralization against some of the most mutated strains of Omicron.
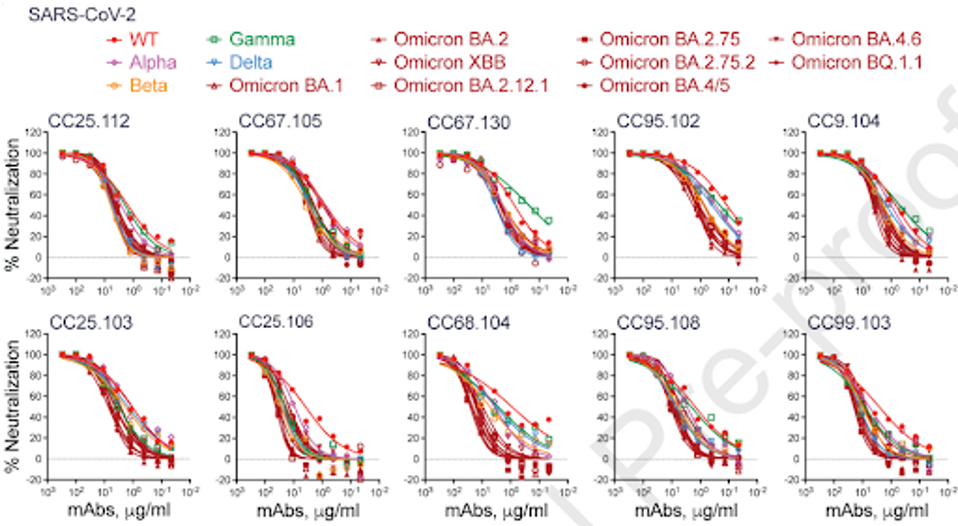
FIGURE 1: Neutralization of SARS-CoV-2 (WT) and major SARS-CoV-2 variants (Alpha, Beta, Gamma, Delta, and Omicron subvariants, BA.1, BA.2, BA.2.12.1, XBB, BA.2.75, BA.2.75.2, BA.4/5, BA.4.6, and BQ.1.1) by 10 select S2 stem-helix bnAbs.
ZHOU ET AL.
Recognition of a Common Hydrophobic Core Epitope
Upon further examination of the 32 broadly neutralizing antibodies, Zhou et al. found that many exhibited dependence on a range of residues on the Spike S2 stem-helix, specifically from positions 1146 to 1156 in SARS-CoV-2.
All 32 antibodies competed strongly for similar S2 stem-helix epitopes, targeting a similar region containing the hydrophobic core residues of the spike fusion machinery. This machinery is highly conserved across betacoronaviruses and may explain why this panel is highly broad in binding and neutralization capabilities.
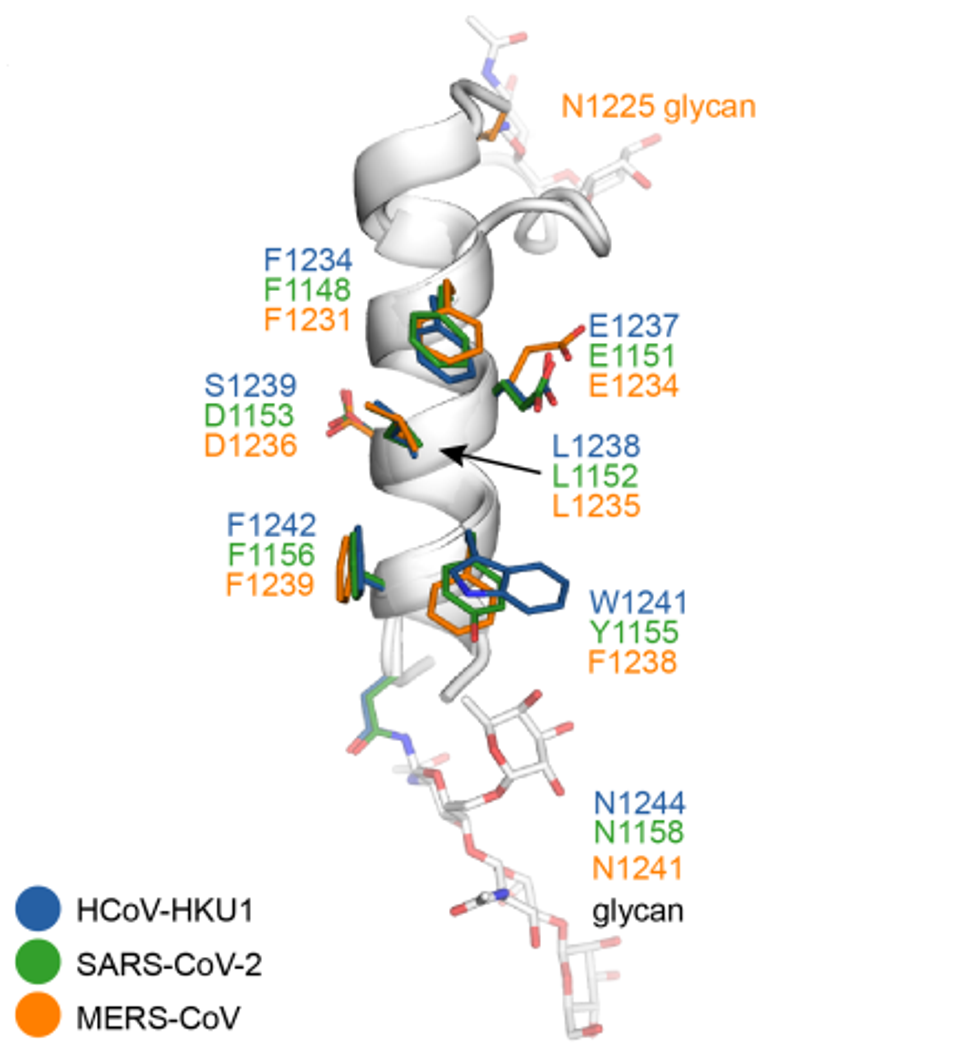
FIGURE 2: Epitope residues in SARS-CoV-2, HCoV-HKU1, and MERS-CoV. Stem helices of these viruses are shown in ribbon representation. Epitope residues involved in interaction with public antibodies are shown as sticks with amino-acid positions labeled
ZHOU ET AL.
Mechanism of Neutralization
To understand the structural basis behind the antibody panels’ broad neutralization, Zhou et al. employed cryo-electron microscopy on four antibodies to determine the complex interactions between the antibody and S2 stem-helix.
The epitope previously described is highly conserved across betacoronaviruses. It is located at the interface within a helix bundle at the base of the prefusion spike. The virus spike undergoes a significant conformational change upon binding with a host cell. Many antibodies aim to block this conformational change to prevent membrane fusion, which is precisely the aim of this antibody panel.
Stem-Helix Antibodies in Live Animal Experiments
To determine the in vivo efficacy of these antibodies matched the in vitro observations, they introduced three of the broadest and most potent antibodies, CC25.106, CC68.109, and CC99.103, to a group of aged mice.
12 groups of ten mice each were given either CC25.106, CC68.109, CC99.103, or a placebo. Each group was then administered mouse-adapted SARS-CoV-2, SARS-CoV-1, or MERS-CoV.
The animals were monitored for daily weight changes, pulmonary function, and viral load in the lungs upon euthanization. Mice that received the antibodies displayed significantly lower weight loss, more normal pulmonary function, and fewer viral titers in lung tissue than in the control group.
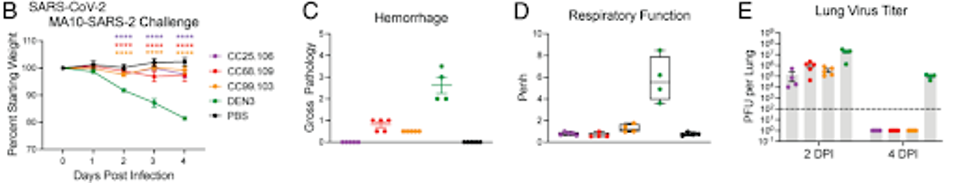
FIGURE 3: (B) Prophylactic treatment of aged mice with S2 stem-helix bnAbs protected against challenge with SARS-CoV-2. (C) Hemorrhaging; (D) respiratory function; (E) lung virus titer.
ZHOU ET AL.
Among the three antibodies, CC25.106 provided superior protection, but all were significantly protective against the three betacoronaviruses.
Discussion
Three years into the pandemic, we desperately need new effective prophylactics and treatments for SARS-CoV-2. Hundreds die from Covid-related symptoms every week in the United States alone. Many of these deaths are in susceptible populations such as the elderly, children, and the immunocompromised, and they could be prevented with an effective and accessible monoclonal antibody treatment.
Unfortunately, many monoclonal antibodies are ineffective against Omicron’s latest variants, as the receptor-binding domain has wildly mutated past recognition by previously promising antibodies.
Enter this panel of antibodies by Zhou et al. Their binding to the S2 stem-helix bypasses the mutated receptor-binding domain for a more conserved virus region. They do this so efficiently that the panel effectively neutralizes even the most mutated versions of Omicron.
We caution that previous attempts at stem-helix antibodies have been made, and they typically lack significant affinity or never made it past initial experiments. That being said, the panel of antibodies by Zhou et al. displays significant promise, and those with the power to do so should pursue them as a potential antibody treatment for the future with haste.


