Making a Chimeric Antigen Receptor
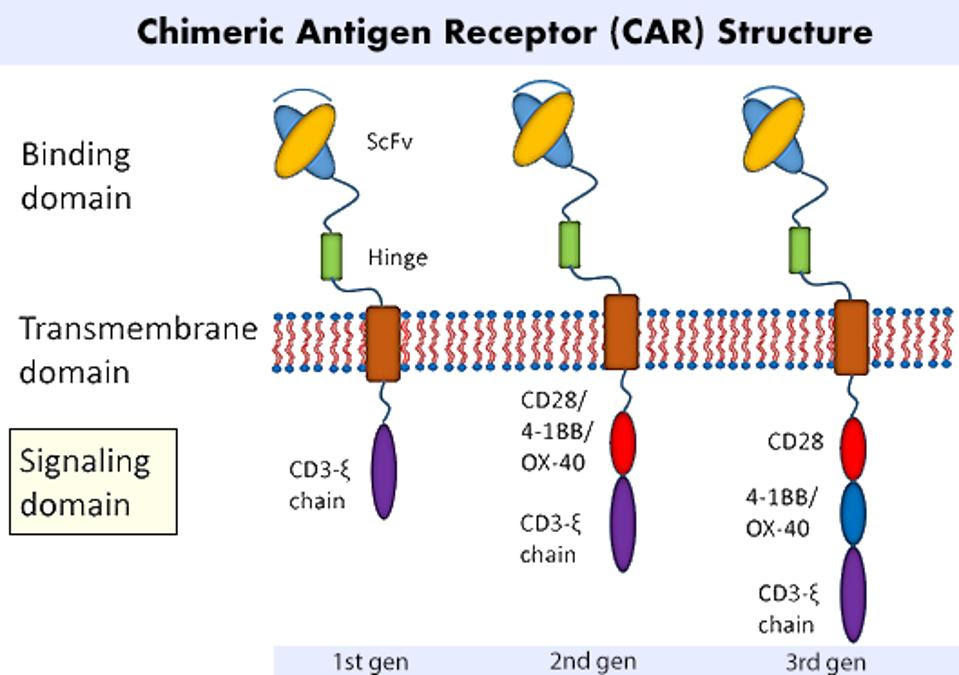
Chimeric Antigen Receptors (CAR) require genetic modification to express new, synthetic components. Figure 1 illustrates the three main regions of a CAR T cell: the antigen-binding domain, the transmembrane domain, and the signaling domain. Scientists often fixate on the binding domain and tailor it for a specific therapeutic target (ex: proteins found on cancer cells). The researchers here, however, focus on the composition of the signaling domain and its influence on CAR T cell performance.
Costimulatory Molecules
The signaling domain of a CAR T cell usually contains a CD3ζ T cell receptor (TCR) molecule and any combination of costimulatory molecules. Costimulatory molecules contain multiple signal motifs, or short peptides which bind to specific downstream signaling molecules. These molecules influence T cell signal transduction to varying effects. Two examples include 4-1BB, which can increase T cell memory and persistence, and CD28, which is associated with effective T cell killing but reduced T cell persistence.
FIGURE 1: A chimeric antigen receptor design can be split into three main regions: an antigen binding domain; a transmembrane domain; and a signaling domain. The signaling domain is most relevant to this study, as the researchers investigate possible combinations of costimulatory molecules to improve the performance and survivability of the CAR T cell.
LIU ET AL. Link Added
Expanding Possibilities with Machine Learning
The researchers at the Wendell Lim lab sought to find unspoken rules governing costimulatory signaling and thereby optimize CAR T cell characteristics. They wielded a synthetic signal motif library, machine learning, and a unique conceptual approach to discover combinations beyond what naturally occurs.
From Words, to Sentences, to Language
The researchers looked to natural signaling proteins, pulled signal motifs from them, and synthetically assembled combinations of signaling motifs to form unique signaling programs. This approach can be conceptualized as a sentence-building exploration.
Figure 2 illustrates this rearrangement of various “words”—signaling motifs—into distinct “sentences” or signaling programs. To understand and predict the “language” of these combinations, the team then used machine learning algorithms called neural networks to detect the underlying “grammar” of the datasets. This revealed the importance of word order, word meaning, and word combinations in the final product—otherwise reframed as the impact of signal motif identity, function and arrangement on T cell phenotype.
The team curated a library of anti-CD19 CAR T cells with assorted costimulatory domains. Each cell contained either one, two or three signal motifs taken from natural signal proteins(see Figure 2). The team randomly inserted 12 native signal motifs alongside one spacer motif into positions i, j and k to yield a total of 2,379 distinct motif configurations, as seen in Figure 3.
Next, researchers screened random subsets from the library to classify the T cells’ cytotoxicity and ability to proliferate (stemness). This process formed unique and unusual combinations, including combinations comparable to costimulatory molecule 4-1BB (ex: M10-M1-M1-ζ).
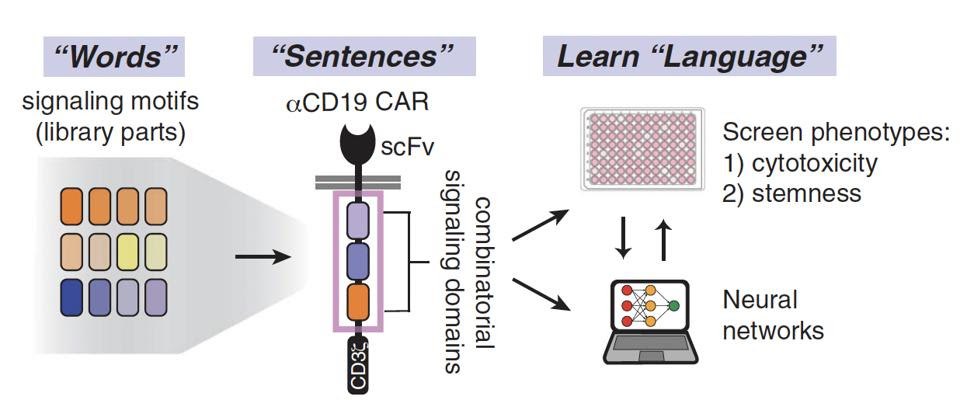
FIGURE 1: The procedure used in this study can be understood through a “language forming” lens. Different combinations of “words,” or signaling motifs from costimulatory molecules, create “sentences.” Machine learning algorithms and neural networks unearth the “grammar” in the sentences—that is to say, the associations between the arrangement of certain signaling domains and the resulting characteristics. Abbreviations: scFv, single-chain variable (antibody) fragment.
DANIELS ET AL., 2022.
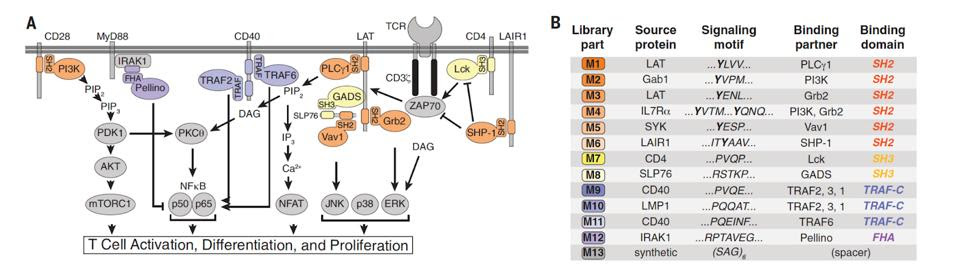
FIGURE 2: (A) Researchers looked to natural signaling proteins such as phospholipase C gamma 1 (PLCg1) to create their combinatorial signal motif library. These proteins play a part in various T cell signaling pathways such as activation, differentiation and proliferation. (B) The library contained 12 native signal motifs and a single spacer motif for a total of 13 motifs.
DANIELS ET AL., 2022.
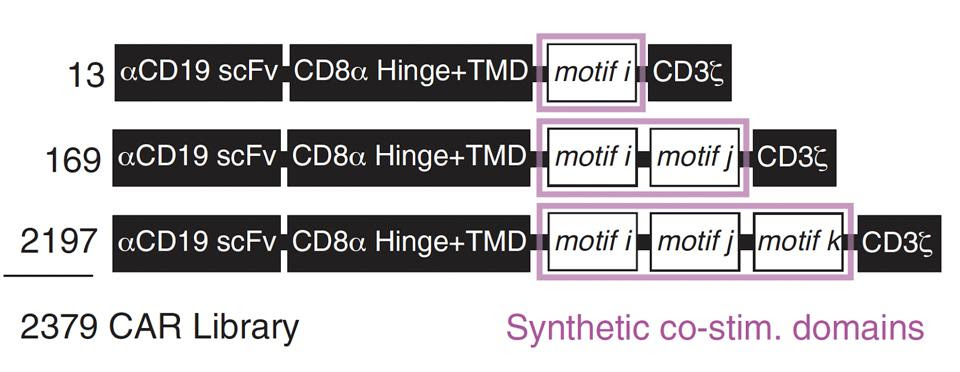
FIGURE 3: The team created a total of 2,379 different motif combinations. All cells equipped the same anti-CD19 antibody fragment and CD8α hinge/transmembrane domain. The signaling domain contained a CD3ζ molecule alongside one, two or three synthetic costimulatory molecules (highlighted in pink). Abbreviations: scFv, single-chain variable (antibody) fragment; TMD, transmembrane domain.
DANIELS ET AL., 2022.
Decoding “Language” Using Predictive Neural Networks
The signaling motif sequences possessed varying levels of cytotoxicity and stemness, according to experimental analysis. The team then leveraged this data to understand the invisible rules surrounding costimulatory molecule design.
An artificial neural network proved crucial to this investigation. As seen in Figure 4, the data was split to either train or test the algorithm to predict a chimeric antigen receptor’s cytotoxicity or stemness. This process elucidated several associations, such as the ability of 4-1BB-like costimulatory domains to enhance cytotoxicity and stemness.
Successful Prediction with M1 Costimulatory Molecule
Could the neural network accurately foretell the fate of a T cell with a particular costimulatory combination? The team tested the waters by adding costimulatory molecule M1 to 4-1BB-like versus CD28-like signaling domains. The neural network predicted that adding M1 motifs would demonstrate enhanced cytotoxicity and stemness in 4-1BB-like domains whilst having no effect in the CD28-like counterpart.
In an in vitro model, the CAR T cells with 4-1BB-like domains and M1 motifs effectively killed tumor cells and maintained T cell stemness; on the other hand, the addition of M1 motifs caused no change for the CD28-like derivatives. This correct prediction translated to mouse model results, as well. The 4-1BB/M1 CAR T cells delayed the growth of tumor cells for two weeks longer than 4-1BB only CAR T cells. These observations demonstrate how a neural network can be used to accurately forecast T cell characteristics depending on involved synthetic signaling motifs.
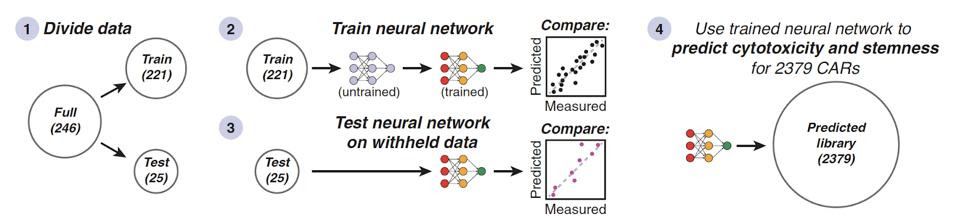
FIGURE 4: The 248 costimulatory molecule combinations were either used to train or test the neural network.
DANIELS ET AL., 2022.
Possibilities for CAR T Therapy
It can be difficult to predict how a synthetic receptor component will influence the resulting cell’s characteristics. This study unscrambles part of this mystery with signal motif libraries and machine learning. By analyzing CAR T cell costimulatory domain combinations, the team created a neural network that successfully predicts T cell phenotype based on the costimulatory molecules present. This, in turn, revealed rules of CAR T costimulatory signaling that can be used to design better synthetic signaling domains. Similar libraries and subsequent analyses could be applied to improve other modular regions of a CAR T cell.